Studies of the genes involved in oil synthesis in microalgae allow scientists to use a gene promoter to increase algal production of triacylglycerols, which in turn enhances potential biofuel yields.
Biofuels are an environmentally-friendly, low-cost and renewable energy source that show promise for reducing reliance on fossil fuels. Although many biofuels have been created from plant and animal fats, scientists are increasingly interested in the potential for large-scale oil synthesis from microalgae.
Now, Hiroyuki Ohta at Tokyo Institute of Technology, together with scientists based at institutions across Tokyo, Japan, have uncovered a way of enhancing the production of fat-based molecules called triacylglycerols in the Nannochloropsis algal strain NIES-2145, thereby increasing oil synthesis from the microalgae.
Triacylglycerols, or TAGs, are a class of lipids which form the backbone to biofuels. The molecules are comprised of glycerol attached to three fatty acid chains, and microalgae is known to produce more TAGs under nutrient stress conditions. When the algal strain Chlamydomonas reinhardtii is starved of phosphorus, TAGs accumulate rapidly following the overexpression of an enzyme known as CrDGTT4, which in turn is triggered by gene promoter SQD2.
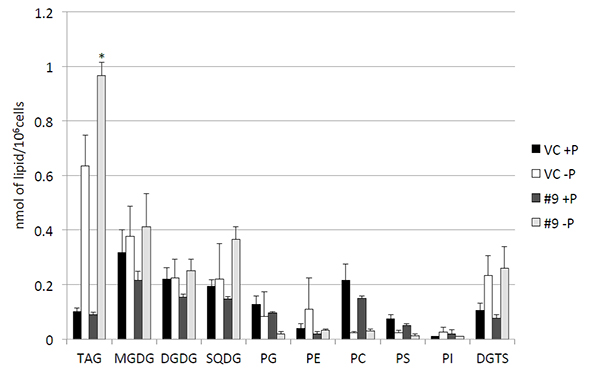
Ohta and his team conducted genetic analysis of NIES-2145 and uncovered a homolog of the SQD2 gene. This implied a common expression control system between algal species in response to nutrient stress. The researchers decided to place both CrDGTT4 from C. reinhardtii and its SQD2 promoter into NIES-2145 to find out if this combination could control levels of TAGs production. Their attempt was successful – the SQD2 promoter was able to drive CrDGTT4 expression in NIES-2145 under phosphorus starvation, crucially without disturbing the membrane structure of the microalgae, and the production of TAGs in NIES-2145 increased as a result. Notably, incorporation of oleic acid (a preferentially utilized substrate by CrDGTT4) into TAG molecules was enhanced.
The findings point to the possibility of manipulating the production of TAGs, and therefore biofuel oil production, in multiple microalgal strains.
Background
Biofuels and how algae could contribute
There are many potential advantages of enhanced biofuel use. The emissions from such fuels are far lower than those of fossil fuels, and crucially biofuels biodegrade much faster — there would be fewer effects on the environment following a biofuel spill, for example. Biofuel is also non-toxic, although one drawback which needs work in fossil fuels is that burning it releases nitrogen oxide into the air, the compound responsible for smog.
The ability to harness the estimated 40-50 thousand litres of oil per hectare per year generated by algae is a prime motivation behind Ohta’s team’s research. Following extensive studies into the model organism for studying algal biofuel production, Chlamydomonas reinhardtii, the team began to find genetic correlations between it and other algal strains. Exploiting these common molecular mechanisms could prove the key to manipulating future biofuel production from algae.
Implications of current research
A key finding of this research is that the SQD2 gene appears common to many types of algae, including primary and secondary endosymbiotic algae. The results of this study indicate that the SQD2 promoter from one type of algae can be used to trigger overexpression of the enzyme CrDGTT4, and thus increase yields of TAGs, in another algal strain. This suggests that genetic manipulation using the SQD2 promoter could increase oil yields from various algae.
Future work
Further investigations are needed in order to fully understand the processes behind lipid remodeling during phosphorus starvation in algae before these methods are trialled on a larger scale.
Reference
Authors:
Masako Iwai, Koichi Hori, Yuko Sasaki-Sekimoto, Mie Shimojima and Hiroyuki Ohta
Title of original paper:
Manipulation of oil synthesis in Nannochloropsis strain NIES-2145 with a phosphorus starvation-inducible promoter from Chlamydomonas reinhardtii
Journal: Frontiers in Microbiology 159028 (2015)
Source
Tokyo Institute of Technology, press release, 2015-09-08.
Supplier
Frontiers in Microbiology
Tokyo Institute of Technology
Share
Renewable Carbon News – Daily Newsletter
Subscribe to our daily email newsletter – the world's leading newsletter on renewable materials and chemicals














